![]()
Cottonwood Ecology Group
Rick Lindroth
University of Wisconsin-
Madison
1630 Linden Drive
Madison, WI 53706
Phone (608) 263-6277
Fax (608) 262-3322
Email Rick
Tom Whitham
Northern Arizona University
PO Box 5640
Flagstaff, AZ 86001
Phone (928) 523-7215
Fax (928) 523-7500
Email Tom
FIBR (Frontiers in Integrative Biological Research Conservation and Restoration)
Community Genetics, Heritability and Evolution: Consequences of Extended Phenotypes:
Deep Background:
Community Genetics:
A recent special feature in Ecology (2003) focused on the
developing field of community genetics (Agrawal 2003, Cavender-Bares &
Wilczek 2003, Chase & Knight 2003, Collins 2003, Morin 2003, Neuhauser
et al. 2003, Ricklefs 2003, Whitham et al. 2003, Wilson & Swenson 2003
), which has been defined as "the analysis of evolutionary genetic
processes that occur among interacting populations in communities" (Antonovics
1992, 2003). In his review of this special feature, Mitton (2003) stated
"An imminent paper in Ecology (Whitham et al. 2003) may herald a new
era in evolutionary biology, the elaboration of community and ecosystem
genetics. If this comes about, evolutionary biology and ecology will be
more tightly linked than ever before." Community genetics is a
frontier in biology because it provides an evolutionary framework for
understanding communities and ecosystems, and it has major conservation
implications. It recognizes that individual species are embedded in a
matrix of 100s of interacting species and that the sum of their
genetic-based interactions has community and ecosystem consequences.
This discipline requires an integrative, interdisciplinary effort that
has not yet been attempted, let alone achieved. Here, we propose a
research program involving genetics, genomics, aquatic and terrestrial
ecology, phytochemistry, biogeochemistry, theory, bio-informatics,
spanning microbes to vertebrates to investigate how genes and their
extended phenotypes affect community structure and ecosystem processes
at multiple levels.
Central to our proposed studies is the concept of the "extended phenotype." Our studies show that mapped genes of a dominant tree produce extended phenotypes in the community and ecosystem, often mediated through plant chemistry. For example, using experimental crosses of known pedigree, we found that a single QTL (Quantitative Trait Locus) accounts for a significant portion of the phenotypic variation in the production of condensed tannins in cottonwood leaves ( Fig. 1A ). Using these same cross types, Driebe & Whitham (2000) showed that the "traditional phenotype" of the concentration of condensed tannins varied ~4X among Populus fremontii , P. angustifolia , and their F 1 and backcross hybrids ( Fig. 1B ). These phenotypes have "extended phenotypes" that go beyond the individual and population to affect community and ecosystem-level processes (Whitham et al. 2003). Because condensed tannins slow decomposition, tannin concentrations explained 63% of the variation in litter decomposition in an aquatic system ( Fig. 1C ; Driebe & Whitham 2000) and 65% of the variation in net nitrogen (N) mineralization in the soil ( Fig. 1D ; Schweitzer et al. 2004). Condensed tannins also affect the foraging of beavers, an ecosystem engineer. Beavers avoid felling genotypes high in condensed tannins ( Fig. 1E ), which then affect the rest of the community (Bailey et al. in review). Furthermore, the selective felling of trees by beaver altered the tannin concentration of the surviving trees; i.e., after 5 yrs, selective herbivory resulted in a 3-6X increase in cottonwood genotypes high in condensed tannins ( Fig. 1F; Bailey et al. unpub. data). These studies demonstrate direct links between a mapped trait and an ecosystem engineer whose selective foraging feeds back to the survival of different tree genotypes. These findings have immense implications for cottonwoods and the thousands of species that depend upon them for their survival. Such extended effects of genes are also important to our emerging national policy on genetically modified organisms due to the potential for transgenic organisms to affect communities and ecosystems (e.g., Strauss 2000, Whitham et al. 2003, Raffa 2004). Using the concept of an extended phenotype, we propose to test three interrelated hypotheses: 1) Biodiversity is an emergent property of extended phenotypes, 2) Extended phenotypes are heritable, and 3) Feedback loops drive community evolution.
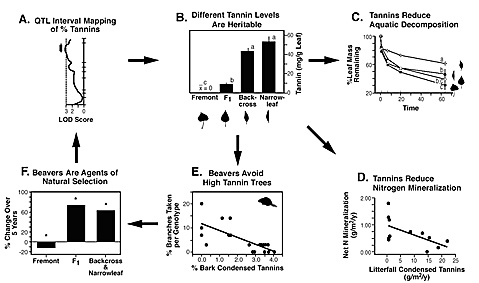
Figure 1
Cottonwoods as a model system for testing these hypotheses - We believe our group can critically address these hypotheses for five major reasons. 1) Our long-term, interdisciplinary collaborations demonstrate that extended phenotypes exist (e.g., Fig. 1). The successful isolation of these extended phenotypes is the first step for testing community heritability and evolution. 2) We have established common gardens with thousands of trees resulting from controlled crosses of known pedigree to test our hypotheses. In addition, we have begun planting three new experimental forests that are important to our proposed research. 3) Because cottonwoods are the first tree genome to be sequenced ( http://www.ornl.gov/sci/ipgc/), the genetic linkages between cottonwoods and their dependent community members can now be investigated. It is important to begin the search for community heritability and evolution with dominant plant species that are known to be community drivers. Our studies demonstrate that genetic variation in cottonwoods is the single most important factor affecting the arthropod community (Wimp et al. 2004) and N cycling (Schweitzer et al. 2004). 4) Our research team has been funded to establish an Environmental Genetics & Genomics facility. This high throughput facility will allow us to process the large sample sizes necessary for examining genetic covariances among interacting species. 5) The development of a long-term parallel experimental system in Australia with Eucalyptus communities enables us to test the generality of our findings (e.g., Whitham et al. 1991, 1994, 1999, Dungey et al. 2000, Lawrence et al. 2003). In each of the following sections, we focus on a key experiment and develop at least one major implication. Space limitations prevent us from describing allied issues, but we point out a few so readers may appreciate the logical extensions of our studies.
Intellectual Merits: Community genetics is a frontier in biology because it provides an evolutionary framework for understanding communities and ecosystems. Our studies of cottonwoods show that genes have phenotypes that extend beyond the individual to have community and ecosystem consequences (i.e., extended phenotypes) . Based upon this concept we propose three hypotheses:
1) Biodiversity is an emergent property of extended
phenotypes. Observational studies have demonstrated arthropod
diversity is a function of genetic diversity among trees within a stand.
By manipulating stand genetic diversity in an experimental forest, we
will verify the relationship between community diversity and genetic
diversity in a dominant tree. We will expand our communities to include
decomposers and mycorrhizae in both terrestrial and aquatic habitats.
2) Community structure, biodiversity and ecosystem processes
are heritable. We will quantify "community heritability" using
synthetic crosses of known pedigree and three independent methods:
a) A novel ordination approach that converts
multivariate data for community phenotypes into univariate "traits,"
then evaluates trait variation using standard quantitative genetic
methods. Our resulting estimates of broad sense community heritability
have proven valid in empirical and modeling experiments. b)
We will quantify the tendency for progeny to support the same
communities of organisms as their parents (i.e., narrow sense
heritability). c) Using QTL studies and the cottonwood
genome sequence, we will map key traits (e.g., phytochemistry) and their
extended phenotypes on the community.
3) Ecological feedback loops result in genetic covariance and
community evolution. We define community evolution as the
genetic interactions among species that influence the fitness of
community members, leading to the evolution of distinct community
phenotypes. Community evolution should be detectable in two ways:
a) Feedback loops - we will quantify the fitness
effects of cottonwood extended phenotypes on species within communities,
and show how these effects feed back to influence cottonwood genotypes
possessing those traits. b) Genetic covariance - we
will quantify how genetic variation among populations of keystone
species (e.g., aphids) covaries with the genetic variation in
cottonwoods. Demonstrating heritable community traits, feedback loops
and genetic covariance sets the foundation for investigating community
evolution.
Our ability to address these issues is enhanced by four major
factors: a) Established collaborations among geneticists,
phytochemists, modelers, community and ecosystem ecologists on two
continents. b) Studies in the wild combined with large
experimental forests of known pedigree. c) Cottonwood
is the first tree genome to be sequenced, providing the genetic
background necessary for our analyses, and d) Our new
genomics center that will dramatically facilitate our genetics
capabilities.
Broader Impacts: Extended phenotypes have major implications
for establishing the genetic foundations of communities and ecosystems,
for quantifying the impacts of genetically modified organisms on the
rest of the community, and for making informed decisions about
environmental policy and management. We have a rich tradition in
training graduate and undergraduate students in interdisciplinary
studies and in mentoring minorities. Our outreach through nature centers
reaches 1000s of visitors, and blends science and riparian restoration
in collaborations with the Bureau of Reclamation, Utah Department of
Natural Resources, Ogden Nature Center , the Arboretum at Flagstaff , US
Forest Service, and Native American tribes.

